I recently gave a lightning talk on the topic of creating your own R packages. Whether you’re working individually or on a team, everyone can benefit from creating a package to organize, document, and reuse existing code.
“Packages are the fundamental units of reproducible R code. They include reusable R functions, the documentation that describes how to use them, and sample data.”
Hadley Wickham
YOU SHOULD CREATE A PACKAGE IF YOU…
Want to easily reuse code
- Packages are an efficient way to reuse existing assets, especially for ongoing projects. Our team has streamlined our ongoing, data science projects by creating packages for our data pipeline process, core metric calculations, and key visualizations.
Spend time copying and pasting functions across files
- If you find yourself repeatedly copying and pasting old code across projects and files, or rewriting functions because you can’t find them, then it may be worthwhile to put it into a package. Doing so will save you a ton of time and frustration in the future.
Need to organize your code in a central place
- Packages help to organize collections of functions and data sets. You could arrange your functions into packages by category, e.g. a data viz package, a package for your biology class functions/datasets, etc.
Collaborate with others on ongoing projects
- One of the easiest methods of distributing code and data for others to use is through a package. Teammates can simply install your package!
Want to standardize your code
- Packages allow you to maintain consistency in your team’s code. E.g. create a standard plot theme with your company’s color scheme, fonts, and logo(s), with set sizes for titles and axis labels. Put it into a package, so that graphs created across teammates will have a consistent look and feel.
STEPS TO CREATE A PACKAGE
I created the following tutorial with my own functions (they’re super simple!) to show the steps of creating a package.
For reference, all my code is on Github. For more details on the package development process, please refer to Hadley Wickham’s book on R Packages.
STEP 1
Install the following packages: devtools and roxygen2.
install.packages("devtools")
devtools::install_github("klutometis/roxygen")
- roxygen2 generates the documentation files for a package.
- devtools is needed for the documentation, testing, and sharing stages of package development.
STEP 2
In RStudio, go to: File → New Project → New Directory → R Package
The following dialog will appear.
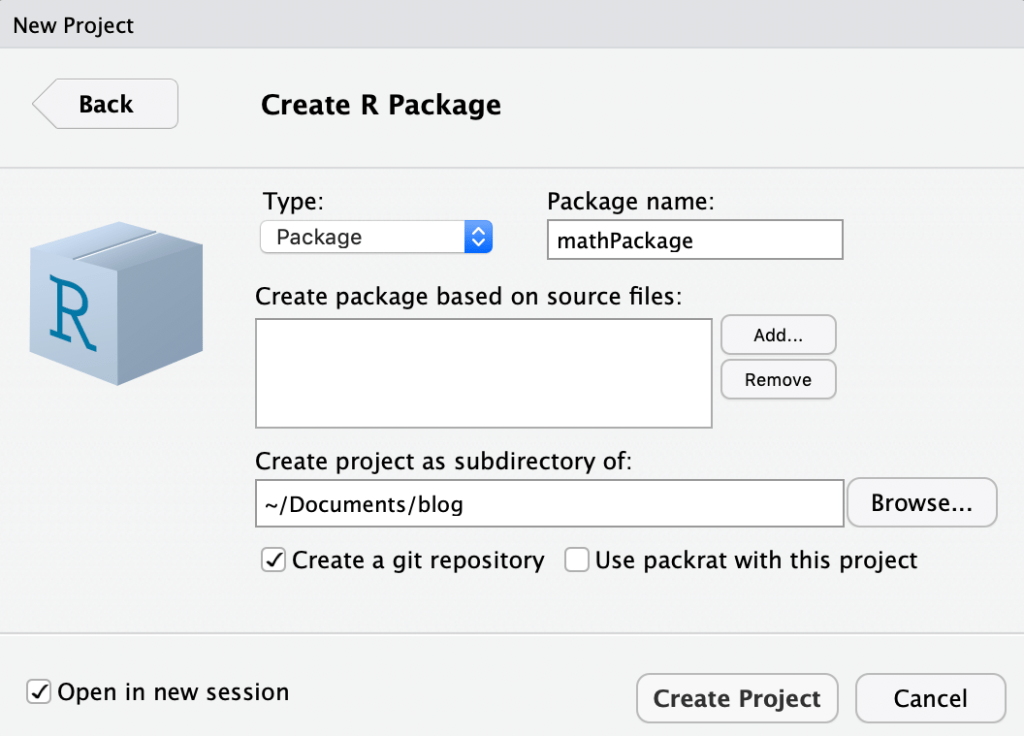
- Enter your package name.
- (Optional) Add any existing R scripts that you’d like to use as the basis for the package.
- Specify your package location.
- I prefer to check the box to create a git repository, for version control. Also, you can check the box to open this package in a new R session (see lower left corner).
- Click “Create Project”. R will automatically build the package, and will create a sample “hello.R” file (which you can delete).
STEP 3
Create a new R script, and start to paste your functions into it. A best practice is to avoid placing all functions in one file, but also to avoid placing each function in a separate file. Ideally, you’ll have a few R scripts, each containing a set of related functions.
See my code for steps 3 and 4 here!
If your code is not in the form of functions, you’ll need to put it into functions in this step.

Save each script to the default location, which will be in a folder called “R”. Packages store all their R scripts in this folder.
Note: If your code uses functions from other packages, these lines of code must be prefixed with the package name, e.g. “lubridate::ymd(…)”. The automated testing that we’ll do in a later step will help you identify which lines are missing a package name.
STEP 4
Now it’s time to add documentation to your functions! This is where the roxygen2 package comes in. Above every function, you will type comments in a special format, which roxygen will later transform into formal documentation.

See this example of how to add roxygen comments to a function (get the code here):

STEP 5
You might be wondering, what happens if my package uses functions from other R packages, but someone doesn’t have those other packages installed? This is where the DESCRIPTION file comes in.
R automatically creates the DESCRIPTION file. See my example file.
On your computer, navigate to your package’s directory, and open the file titled “DESCRIPTION”. This file contains metadata about your package. There are 2 important things to do here:
- If your code calls functions from external packages, list these packages on an “Imports” line (see image below).
- This will cause each package to be automatically installed if someone does not have it installed on their machine.
- Hit Enter at the very end of the DESCRIPTION file. This file must end with a blank new line, otherwise you’ll receive a “incomplete final line found” warning in step 6.

STEP 6
Generate the formal documentation files (.Rd files) by running:
devtools::document()
This enables any user of your package to type ?<function> and see its documentation. Any time you modify the roxygen comments in your R scripts, you’ll need to rerun “devtools::document()”.

STEP 7
Test your package. There are a variety of testing tools, but at a minimum, I like the “check” function from devtools.
devtools::check()
This function thoroughly checks for issues in your code, errors in your package structure, and problems with your documentation.
STEP 8
Use your package! You must first change to the parent directory, as shown in the code below. Install and load your package by running:
setwd("..")
devtools::install("mathPackage")
library("mathPackage")
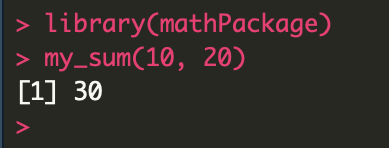
You can upload your package to Github by uploading the entire package folder to a new or existing repo. Then, for anyone to install from Github, they simply need to run the “install_github” function:
# Non-enterprise github
# Modify the "repo" argument if package is in a subdirectory
devtools::install_github(repo = "corinneleopold/packageTutorial/mathPackage")
# Enterprise github - may need to create a personal access token
devtools::install_github(repo = "path-to-package/package-name",
host = "github.hostname.com/api/v3", token = "your-token")
TIPS FOR MAINTAINING YOUR PACKAGE
Whenever you make a change to your package, be sure to…
- Update the roxygen comments (e.g. if you change a function’s parameters or add/delete a function).
- Rerun “devtools::document()” to ensure the man files are up to date, especially if you modified the roxygen comments.
- Commit your changes to Github, then ensure that anyone using your package reinstalls it using the “install_github()” function.
Awesome! This is so helpful.
LikeLike